A New Paradigm for Quantitative Proteomics: Cutting-Edge Mass Spectrometry Solutions Transforming Research
High-quality, scalable experiments increase the value of discovery-phase experiments
For researchers who need to confidently measure small biological changes in their samples, it can be challenging to find a solution that allows them to confidently measure dynamic biology for a wide range of compound classes, let alone for a large number of samples. Complex workflows pose obstacles in scaling up experiments, particularly when conducting targeted quantitation. Find out about various innovations in analytical solutions and the applications being used by researchers today.
What is Proteomic Quantitation?
Proteomic quantitation is the process of measuring the hypothesis-based set of proteins expressed under specific biological conditions. Quantitative proteomics aims to determine the expression level or absolute protein abundance, compare translation levels under different conditions, and identify changes in protein expression that may be associated with diseases, treatments, or other biological processes.
This field has advanced from qualitative techniques to include quantitative analysis using highly sensitive mass spectrometry. This evolution has enabled significant biological insights across various fields like biology, biochemistry, biomarker discovery, and precision medicine. Quantitative proteomics via mass spectrometry (LC-MS) allows for both discovery-based (untargeted) and targeted proteomics applications.
Webinar: Quantitative approaches in discovery proteomics: How do they measure up?
This webinar provides an overview of the current techniques utilized in the field of quantitative discovery proteomics plus a review of the most common workflows and approaches. It also features a study that evaluated the performance of different quantitative approaches using the latest mass spectrometry instrumentation and analysis tools.
Case Study: How mass spectrometry plays a role in understanding the immune system
In this case study, Dr. Susan Klaeger describes how mass spectrometry is one of the few high-throughput technologies that allowed her to measure thousands of peptides presented on human leukocyte antigens.
DownloadApp Note: How to develop and implement LC-MS libraries for clinical studies

Scientists developed a comprehensive metabolite LC-MS/MS library for high-throughput quantitation analysis using the new Thermo Fisher™ Stellar™ mass spectrometer. The new approach showed almost a 3x faster throughput for 2x the number of metabolites compared to previous technologies.
DownloadDeveloping targeted proteomic assays
Dr. Brian Searle’s lab was struggling to analyze low abundance immune cells in the tumor microenvironment (TME) using existing technology. However, with the Stellar mass spectrometer, they were able to confidently profile mouse T cells with reduced starting material because of its high sensitivity and wide dynamic range. Plus – they completed all the experiments in <2 weeks.
Read their paper at the link and watch the video below to hear their thoughts.
Bridging the Gap between Proteomic Discovery and Clinical Testing

Can protein biomarkers be a useful diagnostic or prognostic tool? In a recent study, the Matthias Mann lab used high-throughput absolute quantitation to develop targeted assays for alcohol-related liver disease. They evaluated the Stellar MS for acquisition speed sensitivity to measure targeted peptide panels prepared using a unique labeling strategy.
DownloadHybrid Quadrupole Mass Filter – Radial Ejection Linear Ion Trap and Intelligent Data Acquisition Enable Highly Multiplex Targeted Proteomics
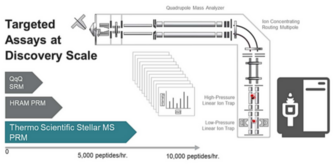
Sometimes the platform of choice comes down to ease of setup and breadth of coverage. The MacCoss Lab at the University of Washington explored how the Stellar MS’ high acquisition rate and intuitive software tools made it an idea platform for targeted proteomic applications. (Spreadsheets and scripting, no longer required!)
DownloadDifferent Proteomic Methods

There are many different methods for analyzing protein dynamics within a cell, tissue, or organism. Which one is best depends on several factors related to specific the research goals, such as sample characteristics and data analysis needs.
For example, isobaric chemical tags, such as TMT tags, are used to quantify relative protein abundance changes in complex protein samples across multiple experimental conditions.