The stages of COVID-19 and its pathology are well characterized, from the contagious phase and symptom onset through to either recovery or, in some cases, death. Briefly, the first one or two days of infection are usually asymptomatic, but the disease is already infectious. The SARS-CoV-2 virus then migrates down the airways, triggering the immune response. For most people, this disease stage is relatively mild and restricted to the upper and conducting airways. However, some individuals experience an abnormal immune response and progress to acute respiratory distress syndrome – a life-threatening condition in which the lungs cannot provide enough oxygen to the body’s vital organs. This exaggerated immune response is associated with the overproduction of cytokines (sometimes called a “cytokine storm”), resulting in dangerous systemic inflammation.
Metabolomic studies can provide insights into these pathobiological processes and, in combination with other cellular and biochemical analyses, can measure the immune response to SARS-CoV-2 infection. Researchers from the Nuclear Magnetic Resonance (NMR) International COVID-19 Research Network have used NMR spectroscopy to establish whether the disease has a metabolic signature at the molecular level by analyzing COVID-19 patients’ plasma metabolic profiles. NMR is ideally suited to this type of research because it enables comprehensive and quantitative investigation of the metabolism and because multiple metabolites can be analyzed in one measurement.

Key research from the Australian National Phenome Center (ANPC) shows a complex pattern of dysregulation of the systemic metabolism caused by SARS-CoV-2 infection (1). The group, led by Jeremy Nicholson, linked this metabolic pattern with multiple organ-specific changes that went beyond primary respiratory symptoms. Similar findings from the Precision Medicine and Metabolism Laboratory at CIC bioGUNE in Spain show considerable changes in the lipoprotein and metabolic profiles of COVID-19 patients – consistent with systemic disease (2). The group described a complex lipoprotein profile in COVID-19 patients and elevated signals for inflammation marker GlycA in the NMR spectra. Using nonlinear statistical models, they separated COVID-19 and non-COVID-19 patients based on these profiles and established a link to increased risk of cardiovascular disease and liver damage in the infected group.
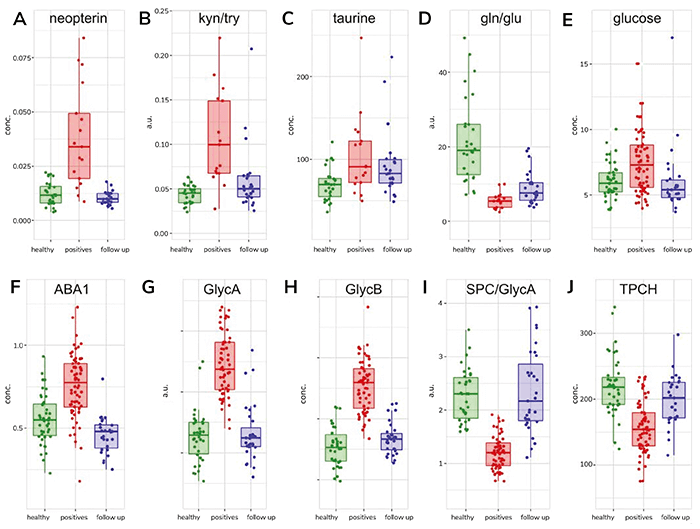
Glutamate plays a major role in cellular energy generation in multiple organ systems, including the kidney, heart, and brain. It is also a key excitatory amino acid in the central nervous system (CNS); elevated levels can induce acute injury. The relationship between glutamate and glutamine is important in immune cell homeostasis, especially for T cells and astrocytes within the CNS, where it acts as an immunomodulator. Additional research is needed to determine the clinical significance of a low glutamate-to-glutamine ratio, but the persistence of a reduced ratio in most PACS patients implies ongoing post-COVID-19 immunometabolic dysregulation.
These results demonstrate variable patterns of functional recovery from COVID-19, with many patients exhibiting residual biomarker signatures months after acute infection. For the first time, researchers have used NMR and MS to measure phenoreversion of some metabolic parameters and report significant metabolic differences between asymptomatic and symptomatic follow-up patients. The Nicholson group’s unique approach in statistically integrating data from NMR and MS into a single dataset using mathematical modeling and machine learning provides a powerful tool for PACS biomarker identification and patient stratification.
References
- VR Anicetti et al., “Immunoassay for the detection of E. coli proteins in recombinant DNA derived human growth hormone,” J Immunol Meth, 91, 213 (1986). PMID: 3525680.
- T Kimhofer et al., “Integrative modeling of quantitative plasma lipoprotein, metabolic, and amino acid data reveals a multiorgan pathological signature of SARS-CoV-2 infection,” J Proteome Res, 19, 4442 (2020). PMID: 32806897.
- C Bruzzone et al., “SARS-CoV-2 infection dysregulates the metabolomic and lipidomic profiles of serum,” iScience, 23, 101645 (2020). PMID: 33043283.
- E Holmes et al., “Incomplete systemic recovery and metabolic phenoreversion in post-acute-phase nonhospitalized COVID-19 patients: implications for assessment of post-acute COVID-19 syndrome,” J Proteome Res, 20, 3315 (2021). PMID: 34009992.
- R Masuda et al., “Integrative modeling of plasma metabolic and lipoprotein biomarkers of SARS-CoV‑2 infection in Spanish and Australian COVID-19 patient cohorts,” J Proteome Res, 20, 4139 (2021). PMID: 34251833.




