Since Jenner first inoculated a young volunteer with his magic cure for smallpox in 1797, the power of vaccination in preventing infection and eradicating infectious diseases has surely been realized. This year, the spotlight has once again turned towards vaccines, as both scientists and the general public cling to somewhat remote hopes of a return to “normal.” Before now, the fastest we have ever managed to produce a vaccine in response to an outbreak was for Ebola – and that took five years to achieve full licensure. The rulebook may have been ripped up, but it is perhaps now more vital than ever that the entire vaccine development process is as efficient, precise, and cost-effective as possible. Developing the right analytical methods using the best tools for the job has an absolutely key role to play.
An analytical chemist with over 14 years’ experience in the pharmaceutical industry, Ewoud started working for Solvay Pharmaceuticals in 2006 after finishing his MBO, but quickly realized he needed at least a Bachelor’s to pursue his dream of becoming an analytical method developer. He decided to take on part-time study alongside his fulltime job, and between 2006 and 2020 he has completed a Bachelor’s, Master’s, and PhD in analytical chemistry while working for both Abbott Healthcare (2006-2010) and Janssen Vaccines and Prevention (2011-2020).
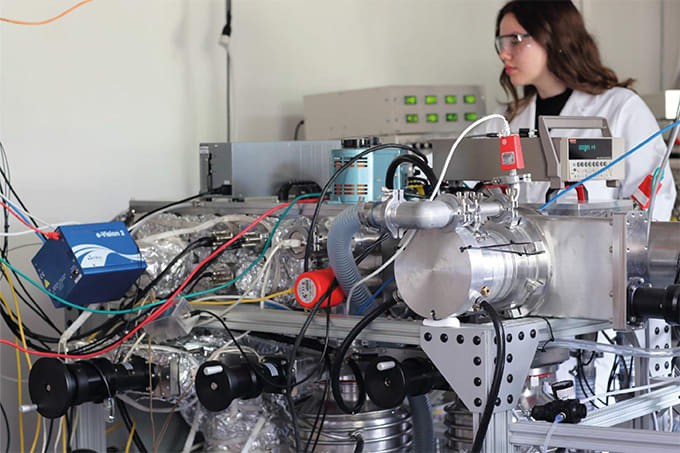
During this time, he’s held many different positions across a broad range of departments. He started working at Janssen Vaccines and Prevention (previously Crucell) as a senior technician in Quality Control back in 2011. Now, he is a senior scientist in the AA department. AA is a group of 25 people within the wider Analytical Development department responsible for the development, validation, and transfer of analytical methods for the analysis of vaccines products. The focus of this group is on developing analytical methods with separation technologies (capillary electrophoresis, LC, MS) and physical characterization technologies (field flow fractionation, analytical ultracentrifugation). Not only do they analyze the main component of vaccine products – typically a virus or a protein – but also the additives and impurities respectively created during the production process or added to the final formulation.
For the last few years Ewoud has been focusing on statistical analysis, dossier writing, and analytical quality by design (AQbD).

Ordinarily, our work at Janssen Vaccines and Prevention – a pharmaceutical company of Johnson and Johnson – is focused on the research and development of vaccine products against infectious diseases like Ebola, HIV, and RSV. So it should come as no surprise that our attention has turned to COVID-19 this past year. Typically, we look at developing modified adenoviruses for intracellular delivery of DNA. Our Advac® technology allows the production of adenovirus vectors in which the viral DNA can be modified to encode an immunogen of interest. In the case of our Ebola vaccine, this is a particular viral glycoprotein. A protective host immune response against the virus is achieved upon vaccination.
The same Advac® technology has been used across our COVID-19, Zika, RSV, and HIV vaccine candidates. Overall, more than 100,000 people have been immunized with vaccines based on Advac® technology, which demonstrates the safety of our platform. Such technology platforms make it possible to quickly develop new candidate vaccines and then produce the optimal ones on a larger scale. Our Zika, RSV, and HIV vaccines are currently in phase 2 or phase 3 clinical trials. On July 22, the first healthy volunteer was injected with our COVID-19 candidate vaccine; interim results from the phase 1/2a clinical study showed that the safety profile and immunogenicity after a single vaccination were supportive of further development. In September, the first patient was dosed in a phase 3 clinical trial to evaluate safety and efficacy of the vaccine in up to 60,000 adults worldwide. In addition to this single-dose regimen ENSEMBLE study, as of November, Janssen initiated a two-dose regimen ENSEMBLE 2 trial which will study the safety and efficacy of the vaccine in a further 30,000 participants. Though we are accelerating vaccine development at the moment, safety and efficacy are never compromised.
My work within the analytical assays (AA) group has been to improve the methods used for analysis of viruses and viral proteins throughout the entire vaccine production process. Our aim was to extend the analytical toolbox for the characterization of vaccine products, helping to overcome the challenges associated with traditional methods. Not only have we developed three new analytical techniques for vaccine development in recent years, we have also implemented a systematic analytical quality by design (AQbD) approach to ensure the right method is developed for the right purpose.
The focus of our AA group is on developing, validating, and transferring methods for different groups within the organization – namely, Process Development, Formulation Development, Product Characterization and the Production Plant. Each department can request method development, validation, and transfer through an analytical target profile (ATP). The purpose of the ATP is to give clear direction to the AA group – it should capture the purpose of the test method, the method requirements, and the reportable results. Importantly, it is defined upfront and agreed between the method developer and the person who requested it.
Clearly, the requirements of any analytical method depend on what it will be used for. For example, quality control methods must be validated, straightforward, robust and reliable. On the other hand, the typical requirements for a process optimization method are a rapid time to result (so as not to delay the production process) and large sample throughput.
We develop analytical methods for a diverse set of purposes:
- Release of material for clinical use – to assure safety and quality of the product
- Stability studies – to assure quality of the product throughout its lifecycle
- Product knowledge – for in-depth characterization
- Process optimization – for example, yield or formulation
After a method has been developed, we then look at its transfer and validation. Analytical methods routinely used for release of clinical material or stability studies will be transferred to the quality control laboratory, which operates under good manufacturing practices (GMP) regulations with validated analytical methods. The methods routinely used for process optimization are transferred to the Biophysics and Process Analytics group, which analyzes up to 30,000 samples per year and is specialized in supporting process and formulation development.
Complex and new analytical methods – or methods that are only used for a single study – are not transferred. In these cases, analysis is performed within our own AA group by those who developed the methods.
Analytical technologies:
- CE (modes: CZE, CGE, cIEF) for virus particle concentration, viral protein content and purity, small molecules, ions (e.g. chloride, Bromide).
- Non-specific colorimetric measurements, such as bicinchoninic acid assay (BCA), Lowry protein assay, and Bradford protein assay, for quickly determining the total amount of protein in vaccine samples.
- Transmission electron microscopy (TEM) to determine the total amount of viruses, including non-infectious viruses, for the quantification of viral proteins.
- LC (modes: SEC, Anion Exchange, HILIC, Reversed phase) for testing of impurities, surfactants, excipients, aggregation.
- AUC (analytical ultracentrifuge) for testing incomplete virus particles.
- MS for protein band assignment after in-gel trypsin digestion.
- FFF (Field flow fractionation) for protein or virus aggregation.
- DLS (dynamic light scattering) for protein aggregation.
Biological technologies:
- Plaque assays for determining the virus concentration as infectious dose.
- TCID50 (50% tissue culture infective dose) for potency – an endpoint dilution assay that is used as an alternative to the plaque assay for viruses that do not form a plaque.
- Single radial immunodiffusion assay (SRID) for the quantification of viral proteins.
- ELISA for protein content.
- qPCR for virus particle concentration and identity.
- SDS-PAGE and Western blot to obtain qualitative data for viral proteins (e.g. molecular weight and identity).
- Agarose gel electrophoresis for DNA content.
The nature of viruses throws up a number of hurdles that must be navigated by analytical method development teams. First of all, the instability of viruses outside the host environment makes it particularly difficult to select the right technology for the quantification of viruses or viral proteins. Viruses are best adapted to surviving and efficiently replicating in the ideal host environment. Outside the host, however, viruses are more easily affected by pH, salt, and temperature changes, which can cause degradation or aggregation. Many technologies require a complex sample treatment to infect cells, to achieve antibody-antigen complexes, to reduce viruses into proteins, or to provide cleanup of the complex sample matrix of viral products (like a vaccine). In addition, many analytical methods require separation conditions – such as organic solvents, surfactants, ion pairing agents or silica-based stationary phases – which may be unfavorable for viruses.
Secondly, the adsorption of viruses and viral proteins can pose a serious challenge; viruses and proteins tend to adsorb to sample vials and instrument parts, such as the injector, valves, tubing, columns, and capillaries.
Finally, the matrix of the crude viral product is typically highly complex and could contain host cell DNA, proteins, cell debris, salts, and surfactants in different ratios and amounts. There is a distinct challenge in separating the virus from these matrix components and preventing their interference with the analytical measurements.
All of these challenges must be carefully considered in analytical method development to ensure successful analysis of viruses and viral proteins.
To overcome the issues typically observed with traditional methods, such as low throughput, limited sensitivity, and matrix incompatibility, our AA team developed three new analytical methods – all previously published – for the analysis of viruses and viral proteins throughout the vaccine production process.
The first is a capillary gel electrophoresis (CGE) method for the quantification of influenza virus proteins and virosomes (virus-like particles) (1). In comparison to single radial immunodiffusion (SRID), RP-HPLC, and SDS-PAGE, the CGE method confers some key advantages. Using the CGE method, we found it was possible to determine three other major proteins in addition to the main influenza protein: HA fragment 2, matrix protein, and nuclear protein. Although CGE could reproducibly separate all four major proteins, quantification was not possible because of the lack of (commercial) reference standards. However, the fingerprint of the CGE electropherogram of the four proteins was specific and could be used to identify the virus strain. The precision and accuracy of CGE was similar to SRID, but the total analysis time for the CGE method was much shorter, allowing analysis of 100 samples in four days instead of ten days for SRID.
The second method we developed uses RP-UHPLC-UV for quantitative adenovirus protein profiling (2). Using our method, all adenovirus proteins could be baseline separated within 17 minutes on a C4 column (300 Å, 1.7 μm, 2.1 x 150 mm) with a water-acetonitrile gradient containing 0.175 percent w/v TFA as the ion-pairing agent. The adenovirus test samples were directly injected into the UHPLC system without the need for sample pre-treatment and the viruses dissociated into the viral proteins upon contact with the acetonitrile/water mobile phase. Our RP-UHPLC-UV method was successfully validated for two purposes: confirmation of the identity of the test sample and detection of protein modifications or degradation products of the adenovirus vector. The method can detect changes in the adenovirus protein composition as a result of thermal or oxidative stress, as well as impurities, such as protein degradants, leachables, and host cell proteins. For RP-UHPLC-UV, the sample throughput was increased by a factor of 6 by reducing the run time from 130 min to 17 min. With the improved run time, up to 50 samples could be run in a single sequence without impacting sample stability.
Thirdly, we also developed a patented (3) capillary zone electrophoresis (CZE) method for precise and accurate analysis of adenovirus samples containing variable amounts of cell debris, cell lysate, host cell proteins, host cell DNA, salts, detergents, and additives (4,5). The CZE method offers an alternative that circumvents issues with current methods – qPCR and anion exchange (AE)-HPLC. Intact adenoviruses from upstream (USP) and downstream processing (DSP) can be directly analyzed by CZE and only samples with high amounts of host cell DNA require a simple benzonase sample pre-treatment. The CZE method has been validated for the quantification of adenovirus throughout the production process. A great advantage of CZE is its compatibility with USP and DSP samples – and their variable matrices. In contrast, AE-HPLC is only suitable for purified adenovirus samples. And with a run time of only 3 min, CZE allows the analysis of 30 samples within 4 hours compared with 3 days by qPCR! Precision and accuracy is also significantly improved compared with AE-HPLC and qPCR. In particular, the improved precision of the CZE method makes it possible to improve the formulation or production process, as smaller process improvements can be detected with adequate statistical confidence.
As part of a continuous improvement project alongside this work, we mapped the process of method development in detail based on input from scientists (6,7). We learned that the complexity of the process and a lack of standardization can result in long lead times for method development and lack of robustness in resulting methods. In short, redevelopment and troubleshooting were too common.
Additionally, for many of the methods the purpose was not clearly defined upfront and that led to improper use or implementation. Analytical method development was typically technology/method-driven rather than product/analyte-driven; methods were often selected because the technique was commonly used, in-house experience was available, or the technique was “at hand.” An assessment to verify whether the selected method is indeed the best choice for the specific product and analyte was mostly lacking. Finally – and especially for complex vaccine products – the matrix and the analyte did not typically match up with the analytical method conditions used.
Put another way, concessions were being made in favor of the analytical technique, but were not optimal for the tested product. The final developed method only produced the “best result” that could be obtained within the restrictions of the analytical method rather than the best result from a given sample. As a result, complex and extensive sample treatments were introduced, and a compromise of suboptimal conditions were being selected.
Based on this information, we decided to implement an analytical quality by design (AQbD) approach when it came to the method development outlined earlier. AQbD consists of six defined steps:
- Definition of the analytical target profile (ATP) describing the objective of the test and the requirements
- Technology selection
- Definition of the critical method parameters by a criticality (risk) assessment
- Method development by design of experiments (DOE)
- Method validation and control strategy
- Method maintenance or method life cycle management
The first challenge we encountered was the lack of guidelines describing the application of AQbD in practice. Typically, only the vision, rationale and a high-level approach to AQbD are described in the literature, meaning tailor-made tools had to be created and developed for most of the AQbD steps. We have now successfully developed and implemented tools for each of the AQbD steps, and we have created training material and courses for scientists in analytical development.
The AQbD process overcomes the issues associated with a lack of standardization. AQbD offers a structured, risk-based approach for method development. The knowledge and decisions made are captured and can be shared and reused. As a result, training of new operators is more focused and there are fewer invalid analyses when the method is applied to real samples.
After all steps of AQbD were applied, and a comparison of six analytical methodologies was carried out, CZE was selected as the method of choice for adenovirus analysis.

Despite the clear advantages, it took years for our CZE method to be implemented within the organization. In particular, we had to overcome prejudice with regards to the robustness of capillary electrophoresis instruments and a general belief that CE could never be run in a QC environment. We sent our colleagues to theoretical CE training to get the background knowledge they needed and our team gave over 50 presentations about the possibilities and versatility of CE. At last, we convinced them (with the data to back it up) that CZE could indeed compete with the current technologies.
Two years after finishing the method development, CZE was to be qualified in a QCD laboratory for their in-process control test of virus particle concentration during the production process. The virus concentration could be reported within 2 hours – it had taken 1-3 days with the previous techniques. An extensive system suitability test and trending of critical data from the analytical method assured them that, after 525 analytical runs (over two years), the precision and bias of the method still adhered to the original requirements from the analytical target profile. Continuous improvement of the CZE test method after implementation and training of the operators proved key to successful daily operation. In 99.4 percent of cases, the sample data could be generated on the same day adhering to ATP requirements. For other techniques, the data was typically reported on the same day in 75–95 percent of cases.
Since then, we have bought eight CE instruments, trained over 20 operators, implemented CZE at six locations, run more than 15,000 samples and we now routinely use 3 CZE applications.
Further to these benefits, the new analytical technologies we developed have also allowed for quick adaptation and implementation with our new COVID-19 vaccine program. I am the responsible scientist for the Ebola vaccine project, but this year I have also been brought in as the subject matter expert for the CZE method that is used for in-process control testing of our COVID-19 vaccine. I am also the responsible scientist for the method used for aggregation determination for characterization of the vaccine product, and have supported the COVID-19 dossier by reviewing the sections describing our analytical release and stability methods.
Our group had two main analytical activities when COVID-19 was announced as our new candidate-vaccine. The main advantage of many of the analytical methods developed in our team is that they can be used for the accurate and precise determination of any type of adenovirus-associated vaccine, such as COVID-19 or Ebola. Our job was to make sure that all these analytical methods were ready to use before COVID-19 production started – thankfully, our platform methods significantly reduce the amount of development and validation work that is needed for a new project. Once all analytical methods used for the characterization of the vaccine were shown to be suitable for our COVID-19 program, they were successfully used to characterize the vaccine batches currently in clinic.
As has been true for many this year, COVID-19 has posed many challenges for us. The characterization of the COVID-19 vaccine is our main focus within the analytical group because a large number of batches are being produced and consequently many COVID-19 samples need to be characterized with our analytical methods. Other challenges that arise dealing with this large workload are the fact that the amount of people allowed in the laboratories is limited due to COVID-19 and this makes it more challenging to deal with the large workload. Furthermore, COVID-19 is just one of our high-priority projects. We also have to deal with the workload for other priority projects that are in phase 2 or phase 3 clinical trials like RSV and HIV.
Besides the characterization of the COVID-19 vaccine, we also spend a lot of resource on transferring our CZE method for IPC testing of the adenovirus-concentration to different production and testing sites, to be sure the IPC-test can be performed on site close to the production plant. Our work typically involves training new operators at different testing sites, but we have not been allowed to visit the testing sites because of the pandemic. Digital training has brought a whole new field of challenges. With more methods needing to be transferred in the future, we expect our workload will only increase, and we are already hiring more staff to try and deal with this.
I am extremely grateful and proud that our AQbD approach has finally offered a standardized approach for method development, validation, and implementation for virus analysis. It’s great to know that our organization is now ready for upcoming guidelines (ICH Q14 and USP <1220>) that will recommend using the AQbD approach. Being able to align different development groups and scientists has fast-tracked our vaccine development program – and we’ve also ensured method development knowledge is captured, reusable, and shareable. Our approach to analysis has not only saved us time and money, it has also provided more information than traditional methods. It has allowed for more efficient production processes, higher quality vaccines, a better understanding of these vaccines, and ultimately made them more affordable.
References
- E van Tricht et al., “New capillary gel electrophoresis method for fast and accurate identification and quantification of multiple viral proteins in influenza vaccines,” Talanta, 144, 1030 (2015). DOI: 10.1016/j.talanta.2015.07.047
- E van Tricht at al., “Fast, selective and quantitative protein profiling of adenovirus-vector based vaccines by ultra-performance liquid chromatography,” J Chromatogr A, 1581–1582, 25 (2018). PMID: 30389208.
- E van Tricht et al., Method for quantification of virus particles using capillary zone electrophoresis (2019). European Patent No. EP 3 259 374 B1.
- E van Tricht et al., “Implementation of at‐line capillary zone electrophoresis for fast and reliable determination of adenovirus concentrations in vaccine manufacturing,” Electrophoresis, 40, 2277 (2019). PMID: 30951206.
- E van Tricht et al., “One single, fast and robust capillary electrophoresis method for the direct quantification of intact adenovirus particles in upstream and downstream processing samples,” Talanta, 166, 8 (2017). DOI: 10.1016/j.talanta.2017.01.013
- E van Tricht et al., “Capillary Electrophoresis in the Early Twenty-First Century: New Trends and Relevant Applications,” Capillary Electrophoresis in the Pharmaceutical and Biotech Industry. Nova: 2019. ISBN: 978-1-53615-223-4.
- E van Tricht, “Advancing virus and viral protein analysis in vaccine development and production: Method development and implementation,” PhD, Vrije Universiteit Amsterdam (2020). ISBN: 9789402819519




