Over the past half-century, advances in the life sciences have been profound, and epigenetics is one of the most exciting new frontiers. The “epi” prefix is from the Greek for “above” - or in this case “in addition to.” It was once thought that the genetic code we were born with was a static blueprint for all that would happen in our cells throughout our lives. Epigenetics describes the revelation that, in fact, that blueprint can be affected to a large degree by the environment. Changes in gene expression dictated by the environment can even be passed down to the next generation and beyond. Many of us would have got a failing grade in high school biology, if we said that information about someone’s life experiences could be passed down to their children. Now, epigenetics is recognized as a basic biological phenomenon.
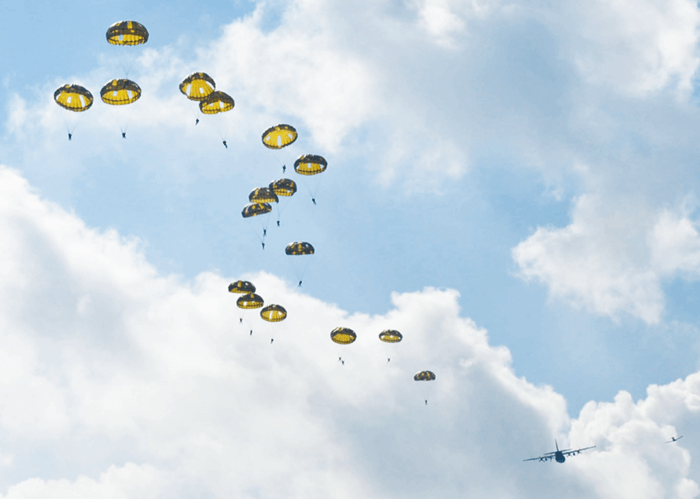
One of the first documented manifestations of epigenetics in humans arose from dire circumstances: war and famine. During World War II, France was liberated after D-day, but the Netherlands remained occupied by the Germans. To liberate the Dutch and hasten the end of the war, British Field Marshall Montgomery developed a plan that became known as Operation Market Garden. The Allies would drop 20,000 paratroopers into the Netherlands to seize a series of bridges along a highway leading to Germany, allowing subsequent Allied troops to move swiftly into Germany.
If you have seen the movie “A Bridge Too Far,” you will already know that the plan was a failure. The Allies encountered heavy resistance from German forces at Arnhem, the bridge over the Rhine River just before the German border. The Allied paratroopers were surrounded by German defenses and sustained heavy casualties, with about 15,000 killed and many more wounded. Even before Operation Market Garden, food was in short supply and, with no reinforcements coming, the surviving Allied paratroopers were soon starving along with the Dutch.
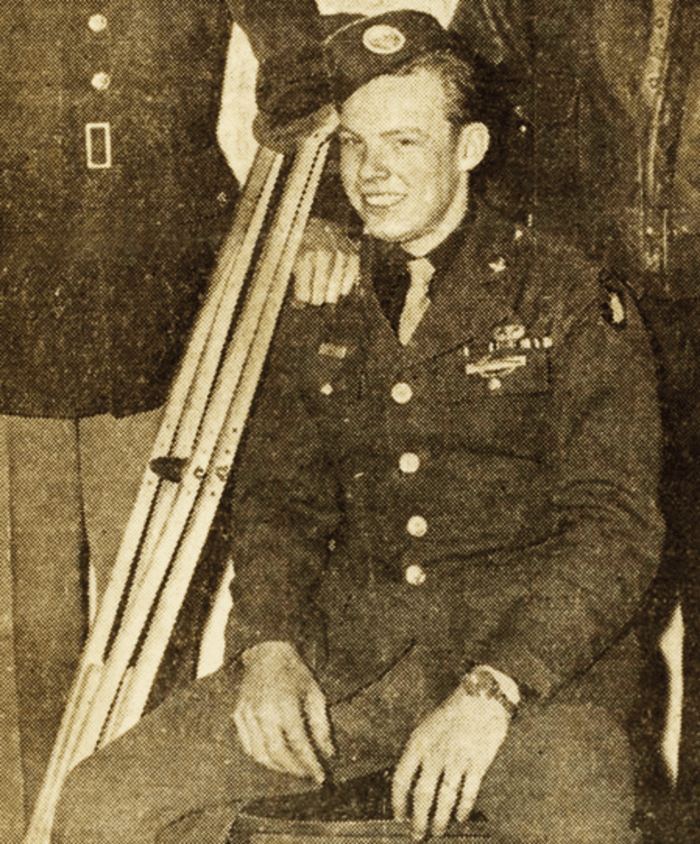


One soldier’s story
One of the paratroopers of the US 101st Airborne, a sharpshooter, flew out of England on September 17, 1944, and his division was dropped over Veghel under enemy fire. His pack was so loaded with supplies that his combat helmet popped off when his parachute opened. But it was easily replaced; so many of his fellow paratroopers were shot as they descended that the ground below was littered with their helmets. His division captured the bridge at Veghel and then marched to Nijmegen, where they helped take control of the bridge over the Waal River. Then came the order to march onward to Arnhem. With German panzer divisions defending Arnhem, and no supplies coming, the Allies ran out of food. They had been told that the Dutch would feed them, and some did, but they had very little food to share, and those who were caught helping the Allies were executed by German soldiers.
One day, the hungry sharpshooter and his best buddy, Bob Sherwood, were out scouting for enemy soldiers when they saw an apple tree laden with fruit. Delighted, they raced to the tree. Bob climbed and started shaking the branches to drop apples down to his friend below.
A single shot rang out.
Bob fell from the tree, killed instantly by a German sniper. Afterwards, the sharpshooter lay on the ground motionless for hours, until he was able to escape under the cover of darkness.
The sharpshooter was not so lucky on October 5, 1944, when he ran in to replace a gunner who was killed by heavy German fire. He knew he would be the next high-value target, and quickly wiped out five gunners with five shots. But a sixth, hidden in the woods, fired a mortar shell that gravely wounded the sharpshooter with shrapnel. He lay in a wine cellar, bleeding, for days before being loaded into a truck overflowing with casualties. A week later, the sharpshooter was transported to the US by hospital ship – and was finally released 10 months later. For his efforts, he earned a Bronze Star for bravery. I am particularly thankful that he survived as he is my father: Robert “Red” Wirth. Now 93 years old, he still tells stories of his harrowing wartime service.
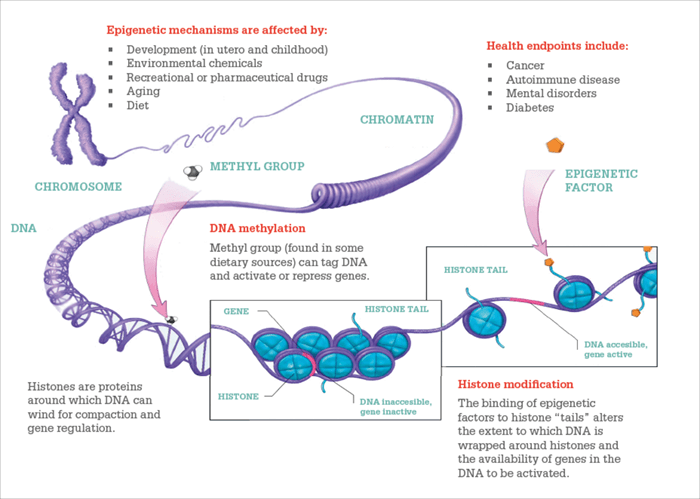
The key mechanisms through which genes can be silenced or amplified by the epigenome.
1. DNA methylation
Genes are described by a sequence of the DNA bases: thymine, adenine, cytosine and guanine (T, A, C, G). A gene is expressed when a transcription factor binds to the promoter region of the gene sequence and commences the construction of mRNA for translation into protein. A gene can be silenced by blocking the binding of the transcription factor through methylation of a cytosine in the promoter region – where the C is followed by a G in the 5’ to 3’ direction (a CG pair or CpG). Gene silencing is nothing new, of course; the cells in your heart contain the entire genetic code, so many non-heart-specific genes must be silenced to allow for specialization. However, it is only relatively recently that we have recognized that environmental factors can silence or de-silence genes through methylation of a cytosine.
2. Histone modification
The other main epigenetic mechanism involves chemical modification of histones in the tail regions, affecting electrostatic interaction with DNA. There are many types of modifications – for example, lysine acetylation removes the charge on lysine to loosen the connection between histones and DNA, for easier gene expression. The various histone modifications are dynamic and reversible, and can work in concert with DNA methylation.
3. Non-coding RNA
A third mechanism occurs later in the process, after the gene is expressed. At this stage non-coding RNA can bind to the mRNA to block its translation into protein.
The thrifty gene
During and after the battle at Arnhem, the Germans destroyed the transportation infrastructure in a bid to secure their hold on the Netherlands, cutting off the food supply just as an unusually hard winter approached. The result was widespread famine. The winter of 1944-45 is known as the Hongerwinter – “the hunger winter”. It was an inadvertent and tragic experiment on the long-term effects of famine. From December 1944 until Germany surrendered in May 1945, food was rationed to under 1,000 dietary calories per day, and dropped as low as 580 calories per day in the height of winter. More than 20,000 people died of starvation.
Dutch babies born early during the famine were born underweight, but after the war, many grew to normal weight. Babies spending only the first trimester in utero during the famine, and born after the war, were born with normal weight. Initially it appeared that this latter group had escaped the worst impact of the famine. However, in the long-term, this group were in fact some of the hardest hit. Far from being cushioned from the effects of the famine, they suffered increased rates of obesity and a host of metabolic diseases affecting cardiac health once they reached middle age. They had somehow acquired what is popularly called the “thrifty gene.” A half-century after World War II, Dutch people born just after the famine found that their bodies behaved as though they were still living under conditions of starvation. We now know that the malnutrition suffered by parents caused epigenetic changes in offspring conceived during the famine.
The conventional understanding of molecular biology was once simple: genes are transcribed to make mRNA, which is translated to proteins:
genes -> mRNA -> proteins.
In reality, things are not so simple – transcription can be blocked by environmental effects, and genes that are supposed to be blocked can be unblocked.
Two main types of gene silencing occurs in cells – one is by modification of the DNA and the other is modification of histones. Figure 4 shows how histones are related to DNA. In the upper left is an X chromosome, and if you started to unravel it, you would find that the DNA is wrapped around histones like pearls on a necklace. The histones are opposite in charge to the DNA to give a strong electrostatic attraction, and modifications can affect how tightly the DNA is held by histones.
Not quite a clean slate
In reproduction, the egg and sperm typically lose their DNA methylation, erasing any memory of the parental environment. The DNA is then re-methylated during development. Folic acid is a source of methyl groups for this critical process, hence the mother’s nutrition in the first trimester is critical. For the Dutch Hongerwinter Cohort, as they are called by researchers, maternal malnutrition meant that methylation was not completely erased, and the epigenetic changes caused by the famine were passed down to their children, and in some cases even their grandchildren – imprinting them for life with the traits needed to survive conditions of starvation.
Scientists in the Netherlands and the USA recently pinpointed the mechanism behind the higher body mass index and elevated serum triglycerides seen in the Dutch Hongerwinter Cohort: DNA methylation at CpG sites in genes mediating energy metabolism (1).
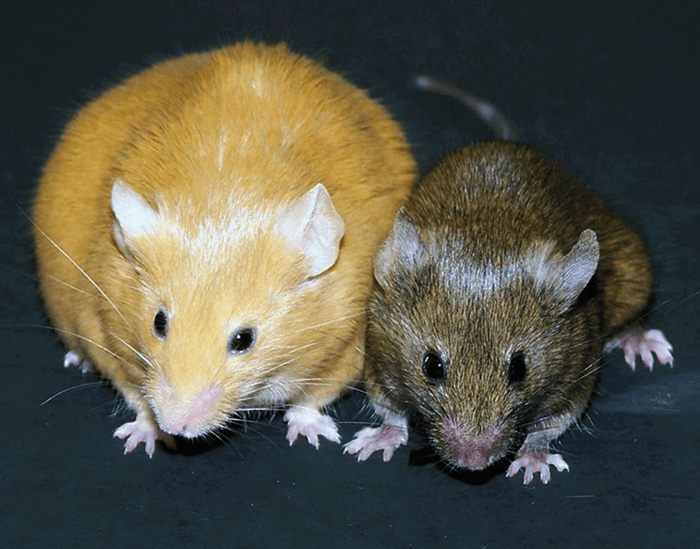
Experiments on animals shed light on the epigenetics of starvation. The two mice pictured in Figure 5 are genetically identical - the only difference is in the diets of the mothers during pregnancy. Both mice were engineered to be pre-disposed for obesity, but the mother of the smaller mouse was fed a much more nutritious diet. Though the genetic blueprint of each mouse is identical, the expression of genes relating to metabolism is radically different, reflected in their disparate appearance. In other words, they have the same genotype but a different phenotype. Hypomethylation of DNA even affects the coat color.
In this case, it was the mother’s diet responsible for passing on unfortunate epigenetic traits, but a study on extreme exercise (mimicking starvation) in male mice showed that the father can pass down the so-called “thrifty” gene or genes through epigenetic changes in sperm, leading to obese offspring (2). However, for women thinking that they would be better off choosing a man who lies on the couch watching TV all day to father their children, too little exercise in male mice causes other epigenetic problems (3).
DNA methylation, once presumed to be a persistent gene silencer, is now appreciated to be a reversible, dynamic process, with an individual’s genome undergoing significant change over their lifespan (4). The average centenarian has significantly less methylation of cytosines than a baby – with de-silencing of certain genes responsible for some age-related conditions (5). Fraga et al showed that identical twins have virtually identical epigenomes at age 3, but by age 50 their epigenomes significantly differ (6), which may help to explain why identical twins usually die of different diseases. For example, the famous twin-sister advice columnists in the US, Abigail Van Buren and Ann Landers, were amazingly identical when they were young, and even pursued the same career path. But they died of completely unrelated diseases, Ann at age 83 from multiple myeloma and Abby at age 94 from complications from Alzheimer’s Disease. As well as this genetic drift, epigenetic changes also lead to predictable effects. The epigenetic clock, for example, is based on a steady rate of DNA demethylation for a specific set of genes as people age – and it is even being used in forensics to estimate the age of crime suspects (7).

Overcoming destiny?
Can we do anything about our aging epigenomes or do we have to make do with what we inherited and put up with whatever changes occur over our lifespan? It appears that we can in fact manage epigenetic changes to some extent. A quick online search for “DNA methylation and exercise”, for example, brings up a wealth of studies. Heart disease is the most common cause of death in industrialized countries, and we know that exercise helps prevent heart problems. Now we know why: our epigenetics change with exercise to help lower our risk (8). And if your genetics pre-dispose you to heart disease, your epigenetics can offset some of the risk.
The breadth of studies on epigenetics is vast. Though we have focused on metabolism thus far, DNA methylation plays a role in a huge assortment of diseases (9). The epigenome of a cancerous cell is very different from that of a healthy cell, and this fact is being exploited for new therapies (10). In other diseases of old age – everything from Parkinson’s disease to chronic obstructive pulmonary disease – you will again find epigenetic factors at work. The diseases that the famous advice column twins succumbed to are both known to be associated with epigenetic changes.
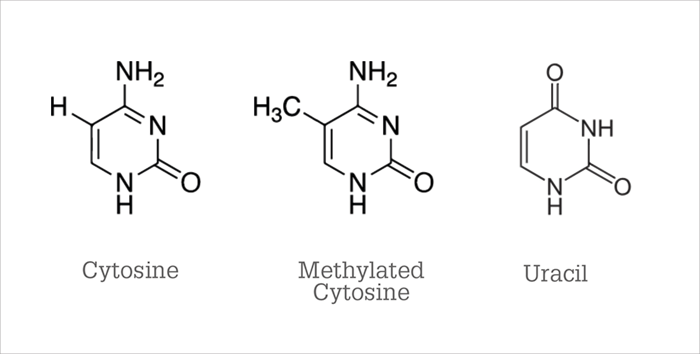
How can analytical science help us to understand - and maybe even control – our epigenome? The key ingredients for a major (or minor) scientific advance are, first, asking the right questions and, second, having the ability to answer them. Analytical chemistry allows us to measure DNA methylation and histone modification to answer the interesting questions raised by biologists. The method presently used for detecting methylation of cytosine was invented in 1992 by Frommer et al, who demonstrated that bisulfite deaminates cytosine, converting it to uracil (11). The structures are shown in Figure 6. Methylated cytosine, also shown, is much slower to react with bisulfite; therefore, methylated cytosines are still visible as C’s in DNA sequencing after treatment with bisulfite.
Analyses in epigenetics began with single-mode measurements; for example, detecting average methylation across all DNA or average modifications across all histones. Once scientists began to understand that DNA modifications and histone modifications work together, new methods were needed (12). The overlapping nature of epigenetic changes poses a measurement challenge, because there is just one copy of each nucleosome per cell. PCR does not amplify methyl modifications, and, of course, protein concentrations cannot be amplified.
As a result of the high sensitivity needed, single-molecule techniques are now widely used for these analyses. In a technique called ChIP-seq, a highly specific antibody for a histone modification (for example, lysine acetylation) selects the nucleosomes with this modification by chromatin immunoprecipitation (ChIP) (13), and single-molecule DNA sequencing using nanopore technology (14) identifies which cytosines in the chromatin are methylated. The nanopore technology sequences DNA and detects modifications without the need for PCR and does not require bisulfite treatment. Future measurement technology will need to address an even larger challenge: there are multiple modifications of histones and other modifications of DNA besides methylation of cytosine at CpG sites. The same histone can have multiple modifications, all of which work in concert with one another and with the DNA modifications. These comprise a complex epigenetic code that describes how our cells operate, how they respond to the environment, and how diseases arise. The limitations of current technology are clear when considering the vast complexity of analytical measurements required to unravel this code (15).
Epigenetics has captured the interest of social scientists, who are concerned about the epigenetics of social status. It is striking that an environment that we have no control over can cause deleterious biological changes, and that this effect can be passed down to the next generation and beyond. The Hongerwinter demonstrated that there can be a critical window, such as the first trimester of gestation, that impacts one’s entire life and sometimes the lives of one’s children. Beyond heart disease, folic acid deficiency during gestation in the Hongerwinter has been associated with a higher level of schizophrenia (16). But malnutrition is just one factor affecting the epigenome - in mice, maternal care in the first months of life has been demonstrated to epigenetically effect stress response in later life (17). Plus, pollution, drug addiction, family dysfunction and stress may all play a part in our individual epigenetic code. These issues and more are discussed in a recent review in the sociology literature (15).
Epigenetics is a rapidly growing and expanding science, encompassing nutrition, exercise, disease, substance abuse, family life and socioeconomic status. There is much we still don’t know, but one thing is certain: progress in all areas would be accelerated with better, faster analytical tools and techniques.
Mary J. Wirth is the W. Brooks Fortune Distinguished Professor – Analytical Chemistry at Purdue University, West Lafayette, Indiana, USA.
References
- EW Tobi et al., Science Advances, 4 (2018). AK Murashov et al., Faseb Journal, 30, 775-784 (2016). KI Stanford et al., Diabetes, 67 (2018). CY Luo et al., Science, 361, 1336-1340 (2018). H Heyn et al., Proc Natl Acad Sci USA, 109, 10522-10527 (2012). MF Fraga et al., Proc Natl Acad Sci USA, 102, 10604-10609 (2005). MJ Jones et al., Aging Cell, 14, 924-932 (2015). T Ronn et al., Plos Genetics, 9 (2013). Q Lu et al., Ageing Research Reviews, 5, 449-467 (2006). MA Dawson, Science, 355, 1147-1152 (2017). M Frommer et al., Proc Natl Acad Sci USA, 89, 1827-1831 (1992). M Berney, JF McGouran, Nature Reviews Chemistry (2018). R Nakato, K Shirahige, Brief Bioinform, 18, 279-290 (2017). AC Rand et al., Nat Methods, 14, 411 (2017). PD Soloway, ACS Chemical Biology, 11, 621-631 (2016). AS Brown, ES Susser, Schizophrenia Bulletin, 34, 1054-1063 (2008). IC Weaver et al., Nature Neuroscience, 7, 847-854 (2004).




