Beer is one of the oldest known beverages – and it remains a firm favorite today. In fact, beer development – and the processes involved – have paved the way for many facets of modern science. An example: fermentation for brewing led to the discovery of microorganismal metabolic regulation through the cultivation of single yeast cells.
Nowadays, food, its safety, and nutritional value are crucial factors regarding our health and well-being. And that’s why we study so-called “dark metabolites.” These are components of food and beverages that are as-of-yet unidentified. In the case of beer, these metabolites persist in spite of the empirical knowledge accumulated through brewing research over the years.
Our mission is to describe the richness of beer compositions – including these hidden metabolites. This knowledge could also enhance our understanding of archeological beer samples and the people who drank them. We’ll toast to that!

Metabolite profiling of food provides extensive and valuable data regarding food safety and quality. Advanced analytical strategies can decipher complex biological and chemical systems and fathom their interaction with each other and our bodies. But, in addition to already well known molecules, a flood of uncharacterized compounds from food and beverages (the dark metabolome!) influence our metabolism and our health. Our work simply stimulates an awareness of that which remains unknown.
Beer just happened to be the perfect matrix for applying our approach. This is in part because of its rich molecular diversity; there is a wide range of raw materials present in beer, as well as additional thousands of compounds produced through processes such as malting, boiling, and fermentation.
We set out to characterize this highly complex system on a compositional level by extracting metabolic profiles, which in turn drive certain attributes of the drink the world loves so much. The aim: to produce a fundamental base of knowledge about the beer metabolome and its origin beyond common databases. Using such a database, old beer and beer-like beverages (archeochemistry) as well as modern industrialized beer (quality control and inspection) can be put into context.
We needed a holistic approach to decipher the diversity, plurality, and complexity of beer samples. But extraction methods and chromatographic pretreatment can limit what can be made analytically visible in terms of polarity and physicochemical properties. Another approach was needed...
A flow-injection analysis (FIA) approach used in clinical metabolomics and for further food samples was our weapon of choice. By diluting beer and then directly and continuously injecting it into the MS-system, we can analyze beer with minimal changes to its chemical composition. With FIA, characterizing much of beer’s molecular diversity becomes a tangible task; however, it’s not all “sunshine and roses.” Such approaches require the highest possible mass resolution to avoid overlapping signals and to differentiate all possible elemental compositions.
The most advanced mass spectrometers in terms of mass resolution and mass accuracy are high magnetic field Fourier transform ion cyclotron resonance (FTICR) instruments. Thus, FIA-FTICR-MS approaches have the power to resolve not hundreds, but tens of thousands of features that might otherwise remain hidden in a very short window of time. The magic happens with a measuring time of 10 minutes and as little as half a drop of beer.
Because of its unmatched mass accuracy, which amounts to 0.1 ppm, it was possible to assign a sum formula and concrete elemental compositions to each mass signal. Or, in layman’s terms, the MS method can assign a compositional name to previously uncharacterized molecules!
But the approach isn’t without its drawbacks... FIA-FTICR-MS lacks information about isomers and concrete molecular structures, which requires a second analytical technique. Another weapon was needed! After raiding our analytical armory for a second time, we decided to characterize the most important molecules on a structural level using UPLC-ToF-MS – and some trusty van Krevelen diagrams!
The van Krevelen diagram makes sense of the compositional information that a molecular composition provides. By plotting the ratio of hydrogen to carbon atoms of a molecular formula against the O/C ratio, we can identify regions in the diagram that reflect the compositional nature of respective molecules and associated biochemical origins.
The tentative classification of metabolite compositions of beer into substance classes lies in their biosynthetic pathway. In gluconeogenesis (glucose production by metabolic processes), the addition of water to the pyruvate gives the carbohydrates very saturated and oxygen-rich compositions, which are located in the upper right region of the diagram.
In contrast, the basic building block of fatty acid synthesis, acetyl-CoA, is obtained via an oxidative decarboxylation of the pyruvate. Another dehydration step during chain expansion leads to less oxygenated lipid species. These can be found on the top left of the van Krevelen. Polyphenols are significantly more unsaturated and have lower H/C ratios.
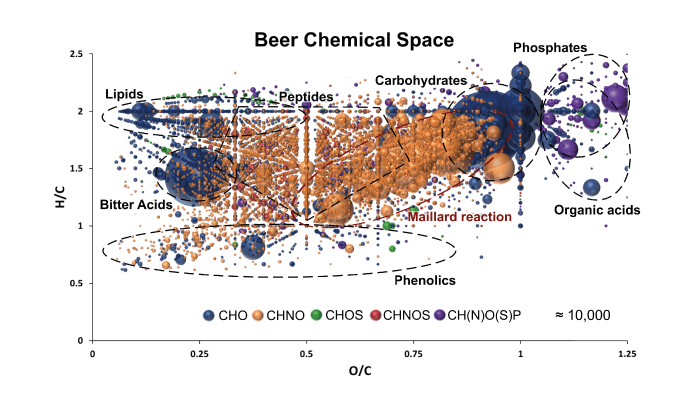
A pint of beer can be mapped according to the corresponding sugar phosphate, nucleotide, and phospholipid spheres. Due to the divergent biosynthetic pathways of the amino acids and the associated different residues, a peptide region is difficult to narrow down. Small organic acids usually have a very high O/C ratio that can exceed the value of one.
Due to their special biosynthesis, the hop-specific “bitter acid” compounds in beer have both the phenolic base structure and the compositional characteristics of terpenes, which the prenyl side chains are based on. Accordingly, these “terpeno-phenolics” show a very characteristic positioning in the van Krevelen. Hence, it is possible to visualize the entire holistic variety and complexity of the beer metabolome in one diagram! But it is necessary and extremely important to say that these classifications are by no means fixed boundaries; they merely represent well-founded reference points!
Our FIA-FTICR-MS approach was able to resolve thousands of yet unknown metabolites in the beer matrix and assign them to possible structural families in the van Krevelen diagram. The definite compositions then enabled us to integrate these molecules into a network via distinct mass differences. These conversions of sum formulae mirror chemical and biochemical reactions and give us information about the processes happening during brewing (and even inside the raw materials used) on a molecular level.
On this basis and through statistical data mining (OPLS-DA), we extracted deep metabolic information reflecting the types of beer analyzed (lager, craft, wheat, abbey). The holistic picture obtained through FIA-FTICR-MS indicated that the hop components in particular differ between these classes. As craft beers are dry hopped (adding hop umbels after the fermentation), these hop components are more oxidized and more phenolic compounds can be extracted.
Far less hops are used when brewing wheat beer, which consequently demonstrates reduced richness of these compounds. On the contrary, the metabolic signature of wheat beer is characterized by the additional grain used; wheat secondary metabolites (phytoanticipines) define the unique metabolic profile of wheat beers. And we were also able to describe a previously unknown derivative of those plant defense molecules, which shows the capability of our approach to investigate hidden metabolites.
Numerous empirical studies suggest that the molecular composition of a beer is very diverse. In brewing literature, rough estimates of the exact number of molecules it contains circulate continuously. We have shown that these estimates are clear underestimations – even without isomeric compounds and molecules inaccessible with ESI and sensitivity limitation.
At last, the typical molecular composition of this unique beverage has been revealed.
This base of knowledge may be used to monitor, control, and guide brewing processes to ensure authenticity, and also to investigate unknown samples (such as historical beers) on a molecular level. By showing the potential of extracting certain molecular patterns from this diversity, our work adds value to modern food safety and quality control. Integrating our foodomic approaches into nutrition and health studies could, for example, help correlate a defined molecular pattern with specific outcomes or observations.
The history of beer accompanies that of our culture and civilization – by no means a run-of-the-mill story of antiquity and tradition. It all started thousands of years ago, when mankind set out to produce durable beverages from domestic cereals. Since then, the tale has evolved alongside jurisprudence (such as the Bavarian Purity Law, which was introduced more than 500 years ago), technology (refrigeration by Linde is a great example), and science (including the discovery of fermentation by Pasteur and single yeast cell isolation by Hansen, as well as today’s cell and gene editing and single-cell metabolomics).
Our study thus provides not only a way to understand beer in previously impossible depth, but also a way to retrace the evolution of brewing means – and even of civilization itself. Different methods of brewing inevitably leave imprints on the metabolic profile of the final (delicious) product. And, for very old but well-preserved samples, it will be possible to trace the methods of brewing used, as well as some of the raw materials and additional processes used.
It may come as no surprise that information of cultural importance can also be extracted from archeological samples with our method. Which type of beer was brewed in a specific region at a specific time? Was it already possible to continuously brew with bottom-fermenting yeast (refrigeration)? Are there any clear indications for or against complying with the purity regulations? How was the grain malted (as is reflected in its darkness and dark metabolome)? Answers to all of these questions can be found within.
Admittedly, samples of beer or beer-like finds from earlier human history are rarely so well preserved that one can work with a liquid that has remained behind. In these cases, the residue crust must be examined, and its molecular pattern compared with the beer we know today. We are not limited to individual potential marker molecules, which often are ambiguous. We can offer whole metabolite profiles from hops, used cereals, yeast and fermentation or Maillard processes that offer an extensive base of knowledge about the beer’s metabolome.
As for the future, we see this study as a starting point. We hope to open and answer many more questions about the brewing industry – and our society – as time progresses. So let’s raise a glass to the future – cheers!

We would like to acknowledge Marianna Lucio and Michael Rychlik Prof for their valuable contributions to this work and Martin Zarnkow and Patrick McGovern for their stimulating discussions in the last decade on beer and civilizations - we couldn’t do it without them!




