For more than five decades, I have worked in academic research. The questions I and my colleagues – graduate students, postdocs, and collaborators – addressed in the beginning were “academic”, meaning that they focused purely on curiosity. They were usually great fun, but often seemed a little other-worldly. More recently, I have become interested in how best to make university research both intellectually interesting (that is, science for the sake of understanding) and practically useful (that is, technology that works). When the two occur together – or at least in the same project – the result is research centered in what is called “Pasteur’s quadrant”: these are problems of major societal importance for which the science base to generate a solution does not yet exist. The problems in Pasteur’s quadrant raise philosophical and practical issues about the value of sophisticated, complex research, relative to research guided by the idea of a simple (functional) solution. My current belief is that while complexity can be beautiful, simplicity works better.
As a scientist, I have become increasingly radicalized with age: science has purpose beyond simply amusing scientists. For many years my research was almost entirely curiosity-driven, but at a particular point – starting roughly with the initial development of self-assembled monolayers (SAMs) – I began to suspect that this style was neither the best possible use of my time, nor the best way to provide a broad education for students. It was no coincidence that the field of surface science that developed around SAMs had a low barrier to entry, and could address both scientific and practical problems efficiently. By combining simplicity with broad experience in multi-functional teams (and, occasionally, start-up companies), our approach to research has gradually become more interventionist – both in terms of science and applications. The range of problems on which we now focus includes the lack of appropriate healthcare technology in the developing world (and the exorbitant cost and ineffective performance of the healthcare system in the United States is a problem that isn’t far behind), the origins of life, the behavior of dissipative systems and the mechanisms of quantum tunneling; a research program that combines “curiosity” and “use.”
As scientists who get our money from the public purse, we have an obligation to spend some time producing science that helps to solve problems. (There are, of course, differences in opinion on what strategies for research best serve the interest of society.) In some cases, rather than developing more and more elaborate extensions of the very sophisticated science that’s already out there, I (and especially my colleagues in the research group) prefer to start with a clean page and ask, “what can we do that solves a problem intellectually or practically, and that’s very simple?”
That’s where I am right now. How I got here is a long and winding story…

When I was young, there was no such thing as a helicopter parent. Children were almost thrown out of the door after they’d been fed breakfast and were expected to come home for dinner when they got hungry. In between, they amused themselves, got into trouble, and explored. Not only was it mostly wonderful fun, it also instilled the idea that the world is full of things that you can’t always understand but that you can tinker with. My father was a chemical engineer and ran a small chemical engineering company in my hometown of Louisville in Kentucky. Despite this connection to chemistry, my route into academic chemistry was based on luck, and, to some extent, coincidence. I hadn’t considered fancy schools on the East Coast as an option (in fact, the idea of “options” was not part of my vocabulary: as most children, I simply went to school), even though I was raised in a middle class family and attended private school. One day, seemingly out of the blue, a teacher from my school called my parents and said, “Can I come out and talk to you about your child?” (a question that usually makes a parent’s heart sink). This teacher – a stranger to me – insisted that my parents send me east to prep school, and had arranged for me to go to Phillips Academy in Andover, Mass – a move that he had organized purely out of the goodness of his heart. It didn’t strike me as being as remarkable back then as it does now. The step up from Kentucky to Andover was a big one; but from there, Harvard, CalTech, and MIT followed quite naturally.
I went to Harvard University in 1957, intending to study English or mathematics, but found that I lacked the talent to be in mathematics. I also couldn’t imagine how I might make a living from English. As a teenager, I had worked at my father’s company over a couple of summer holidays as a technician, so I was reasonably experienced at routine laboratory work. I liked working with my hands, washing dirty dishes, doing distillations and so on, so chemistry seemed like an effortless choice. It was not a deeply analytical and thoughtful decision on my part. Still, to be a good fit for a profession you need to enjoy doing the day-by-day stuff, rather than being focused on some ultimate (and usually not well-imagined) view of success. Since I was at least an experienced technician, I did well enough, and chose to stay in Chemistry. Chemistry at graduate school was a logical progression, and from there I went to work in a university, because it seemed more appealing to work on whatever I wanted to work on, rather than doing what somebody else wanted me to do.
My PhD work, in the early 1960s, was done with a distinguished physical organic chemist named Jack Roberts at the California Institute of Technology (“CalTech”). A big component of Jack’s work was applying the tools of physical chemistry to problems in organic chemistry, and one of the hot instrumental techniques at the time was nuclear magnetic resonance (NMR) spectroscopy. By today’s standards, it was unbelievably primitive: you didn’t need to scan the NMR spectrometer in any direction because the drift in the field was so rapid it would just scan itself randomly up-and-down and back-and-forth – you’d get half the spectrum of ethanol and, later, the other half of it backwards, all of which was pretty amusing. Thankfully, we were not analytical chemists; our interest was in structure and mechanism. By the mid-1960s, I had settled into the Massachusetts Institute of Technology and was exploring an interest in polymers, which I knew very little about. MIT was, and still is, a wonderful place to be because of the diversity of interests and experience to be found there. Around that time, the first steps towards biotechnology were being taken (it was called bioengineering then) and Ray Baddour, the chairman of Chemical Engineering, was putting together a team for a bioengineering project. In such a team, you need one of everything: a fermentation engineer, a geneticist, a fluid mechanic, a microbiologist, and so on. The team was all set and ready to go when the chemist, who was an expert on enzymes, decided to move to Israel, leaving a gap. Ray started with the letter A in the list of chemistry faculty at MIT and called everybody in alphabetical order, asking, “Would you be interested in helping us to figure out what bioengineering is?” And everybody, very sensibly, said, “no”. Until he got to W for Whitesides, at which point I said, also very sensibly, “yes”. In general, I think it’s a better answer than “no,” and knowing nothing about polymers was intellectually very similar to knowing nothing about enzymes. I was, however, doing a lot of work in organoplatinum chemistry and part of the reason to be interested in biology was because platinum is such a good catalyst – and, of course, the same is true of enzymes.


The subsequent four years were fantastic. I worked on biology, applied biology and bioengineering – fields that had previously been completely unknown to me. One lesson from this experience that has really stuck with me is that it’s okay, if you don’t know anything. Just jump off the end of the springboard and see if there’s water in the pool! You’ll win more often than you lose, and if there’s no water, well... I discovered that there are things you can do with enzymes that you can’t do with ordinary catalysts. Interestingly, when I suggested to the superb synthetic chemists around me that they might try an enzyme in place of platinum or sulfuric acid catalysts, the idea was met with horror. Regardless, and insensitive to this most unenthusiastic reception, I set off down that path, with chiral compounds and sugars being the two main areas of focus. I never thought of it as applied research, more as a program in exploratory catalysis.
Catalysis – at least the large-scale heterogeneous catalysis practiced in the chemical industry – is, of course, all about surface chemistry. At that time, everybody in surface chemistry was working with beautifully clean single crystals of, for example, nickel, prepared in ultra-high vacuum, and studying reactions on these beautifully prepared surfaces, using sophisticated instruments to do different kinds of spectroscopy. To me, it all seemed just too complicated, slow and expensive, and I wondered if we couldn’t come up with a form of surface chemistry that was a lot easier to do and also more relevant to biology and organic chemistry. We started with polyethylene, which is the prototypical organic polymer, and found that it could be oxidized to introduce carboxylic acids onto the surface; organic chemists like carboxylic acids because you can do a lot of different kinds of chemistry with them. This was in the eighties and, now back at Harvard, I worked on the topic for a decade, a progression that took me from catalysis to heterogeneous catalysis, to surfaces, to organic surfaces, to polyethylene.
My interest in self-assembly once again came from the notion of doing something that no one else was doing. Organic chemistry was dominated by extraordinarily complicated, interesting, sophisticated synthesis, of mostly natural products. The reactions were really beautiful constructs of science and, because they involved making covalent bonds, were based on making non-equilibrium structures. By this time I was a functional biochemist, and realized that most of the important structures in biology – lipid bilayers, folded proteins, paired strands of DNA – are not made entirely out of covalent bonds. Amino acids are linked together covalently, but that string is completely inactive until it is folded; a bilayer lipid membrane isn’t stitched together, it just forms itself like a soap bubble. Everything in biology basically puts itself together. I became fascinated by the idea of using systems that put themselves together rather than relying on bond-after-bond synthesis by chemists. Self-assembly is the answer (or one answer) because it creates molecular aggregates that are very much bigger than those that can be made with conventional synthesis: crystals, protein-like structures, films on surfaces, all kinds of stuff.
Our focus was on developing new methods of making large molecular ensembles or ways of mimicking biology – but these were exploratory studies that were not directed towards solving a specific problem. Nevertheless, the invention SAMs was really a big deal for us (and for surface science more broadly). Our work started in a joint activity with Ralph Nuzzo, a distinguished surface and material scientist, now at the University of Illinois. Ralph had been a student with me, working on organoplatinum chemistry, and had gone off to work in Bell Laboratories. He came by the lab one day, and very generously told us about a process he and Dave Allara were using to create very well-ordered SAMs on evaporated gold films. We started to work together, with Ralph providing the crucial spectroscopy and with us doing the physical organic chemistry to show how you went from molecules to ensembles of molecules with material properties.
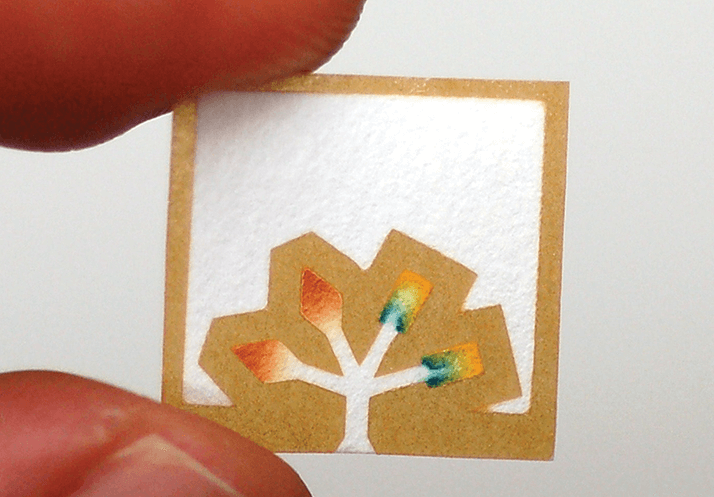
The idea of simplicity was a very powerful concept that I took, in part, from our work on self-assembly. It was, for example, straight-forward to print SAMs, which got us into something that looked like lithography. Then we realized that by sealing the elastomeric stamps we used for printing to the surface (instead of using them to transfer the “ink” that became the patterned SAM), they became microfluidic systems. This led to a ten-year run in which we – along with Andreas Manz at the Institute for Advanced Studies in Freiburg, Germany, and Steve Quake at Caltech and Stanford – invented much of the foundation of what’s known as microfluidics. Paper diagnostics – the ultimate in simplicity – was the next step in the chain. While we had a versatile fluid-handling device that offered a relatively inexpensive way of doing analysis, it was by no means cheap. One of my post-docs, Sam Sia, was very interested in using microfluidic systems to create analytical devices that would be inexpensive enough for the developing world. He introduced us to the subject and his devices worked well, but I became interested in making simpler and even less expensive solutions. Certainly, the diagnostics for the developing world was another driver of this interest in simplicity, but so was the idea of more broadly-applied health surveillance in the USA. A further motivation for paper diagnostics was our desire to undertake another kind of experiment: to determine if it is it possible for a university research group to create a new technology to be used in the field by real people.
This was a step into the unknown of a different kind – one no longer restricted to research questions, but encompassing societal, political, regulatory, and financial issues. One wonderful thing about being a scientist is that, within reason, you get to do anything that you want, and very smart people will often help you learn. I had been involved in projects with the Defense Advanced Research Projects Agency (DARPA) and with a number of other advisory groups in Washington, DC. This work was very instructive, teaching me something about technology and about public policy. I learned all kinds of things that then became part of how I think about science. One of the consequences of this broadened education has been the development of Diagnostics For All (DFA, see sidebar), a not-for-profit company that makes analytical devices for biomedical use in the developing world. Our laboratory at Harvard now works cooperatively with DFA to invent, prototype, engineer and validate low cost biomedical devices.
To me, an analytical instrument is simply something that generates information. The big change in analytics is the migration from generating analytical data to generating actionable, affordable information. I attended Pittcon this year and the exhibition contained endless booths full of fascinating devices, and most interesting solutions to problems in analysis; some of them – but only some – also considered their product as information. More focus on information in biomedicine could perhaps help us to think beyond individual health, and beyond those diseases where all that can be done is to put off the inevitable temporarily. Instead, we could think about public health, about managing cities, about understanding the environment, about food security and water safety. To measure these complex systems requires lots of data, so data acquisition has to be cheap. And, although there are some things where critical measurements require laser spectroscopy or some other highly sophisticated instrument, anything to do with food, water, your internal plumbing, or the safety of animals in South Africa also requires taking samples of fluid, be it blood, serum, tears or sweat. That’s where wet (bio)chemistry is always going to come into it. Our focus has been on using technology to make things fundamentally cheap, and ideally convertible into smaller pieces that can go into the analytical web.
The American medical system works in a for-profit mode; for example, if I get a prostate-specific antigen (PSA) measurement it might cost fifty bucks. Every other high-technology industry has made the transition from “analysis for profit” to “information for free” that medicine has not made. For most high-technology areas, measurement has ceased to be a profit opportunity. Instead, information is free, and the value is added by those who manipulate it, store it and distribute it. To give an example, the number of bits required to fly an airplane is simply staggering, from how you set the ailerons, to how the aircraft is tracked by air-traffic control. Yet the pilot doesn’t come out of the cockpit and get you to drop five dollars in quarters into a slot beside the wing every time he or she wants to reset the ailerons. But that’s basically the way the medical system works – every time data is needed, somebody gets paid for it. I believe that’s simply the wrong approach, so part of our interest in paper diagnostics is to create a situation where information in medicine is as close to free as possible. The value should then be added by doing other things with that data. You see this concept in spades in genomics. The cost of sequencing has fallen at an astonishing rate – faster than electronics. It is now possible to sequence an entire human genome for $1,000, and the cost will continue to fall. Soon, every baby will start life with a full sequence on a USB drive. That economy of scale – presently made viable by the functional simplicity of Illumina sequencers – makes it possible to think about medicine in fundamentally new ways.
Looking at the American business model in healthcare, it is hard to say how much of what gets done is for patient benefit and how much is for profit. Instead, we should think about information in the biomedical system as being a public good at lowest possible cost. Don’t misunderstand, I’m a hardcore capitalist, but I think that the reason that society tolerates the entity called a corporation is both because it generates a return on investment and because it provides solutions to societal problems. One has to be a little careful that the tail does not wag the dog. A wonderful thing about universities is that if we get interested in bringing high technology to public health – as opposed to bringing it to end-of-life medicine – and if we can find somebody who will pay for it, we can do it. That, in my view, is exactly what a university should be doing. We should be doing things that are not accepted as a proven paradigm in society. Ideally, we should be doing them simply so that other people who are interested can get involved and take those that turn out to be really good ideas, and expand them to the benefit of society. Do I think that it is important for scientists to do things that have societal benefit in mind? Yes.
Over the course of my career, I have been involved in a dozen start-ups in different areas. The process from invention to product (and by a product I mean something that solves a specific problem that’s sufficiently valuable that somebody will pay you for it) is not a simple one. It sometimes (but not always) starts with a new idea; then you have to show that the idea actually works and that it might be good for something; then you have to make a crude prototype of the product and somebody has to understand that it could be adapted to solve a real problem; and then they have to make a prototype of the real thing that someone might want to manufacture. Manufacturing is another art form altogether, requiring lots and lots of money, regulatory clearance and so on. You have to think about distribution and training, advertising and sales, all kinds of things. The process requires cooperation among a series of entities, ranging from the university to a pharmacy chain like CVS. A really interesting intellectual problem is how you put together these entities. One of the things that I have concluded is that it is very difficult to run a single research organization in which part is tasked with creating new things and part is tasked with doing engineering development. Engineers are clever at taking something that exists and making it better, cheaper and faster, but they need some starting point; scientists are good at imagining and inventing things but not at development. It’s not that engineering is more or less creative than science, it’s just that engineering and science are creative in different ways, and each requires people who enjoy the particular set of activities required for their part of the problem. Given all this, Diagnostics for All (DFA) was launched as a not-for-profit company explicitly to do product development based on the inventions from my lab at Harvard in the paper diagnostics area. Today, about five years, DFA’s first product has undergone successful field trials in Africa and Vietnam, and several other technologies – for immunodiagnosis and nucleic acid analysis – are at advanced stages. Meanwhile, in the research lab we are developing hand-held electrochemistry and two-phase polymer systems. It is a very successful collaboration in which we, Harvard (and any other relevant entity – technology should be agnostic on the subject of parenthood), invent, and they, DFA, look at problems and engineer solutions. When involved in start-up companies as a university scientist, you have to work with them in those periods when you can contribute (there are times that they don’t actually need – in fact, don’t like – outsiders very much). At the moment, DFA is something that I’m spending a lot of time on, because I believe that I can make a contribution. With for-profit start-ups there is a clear goal, which is to make money and to pay back the investors. With not-for-profits the story is a little bit more complicated because you have been given money with the expectation of contributing to societal good. I can’t say which I prefer, that’s like being asked which of your children you prefer. But doing something that’s very interesting, possibly useful, and new is always satisfying.